Our group also works on many programs to address important developmental questions, such as sex determination mechanisms and limb development in vertebrates
- Genomic mechanisms of sex determination in vertebrates
The existence of males and females, which are often strikingly different in morphology, reproductive strategies and behavior, is one of the most widespread phenomena in biology. However, the genetic mechanisms that generate this ubiquitous pattern are surprisingly diverse and do not follow a phylogenetic pattern. Vertebrate species possess diverse sex determination mechanisms, eg. Mammals have male heterogametic sex chromosomes (XX in female, XY in male), whereas birds have female heterogametic sex chromosomes (ZW in female and ZZ in male). The sex determination in other vertebrate groups like fishes, amphibians, and reptiles, is controlled by a variety of mechanisms, including sex chromosome determination (XY or ZW), sex loci determination, or environmental sex determination. Our research interests on sex determination are on the evolutionary process of the sex chromosome pairs and epigenetic regulation mechanisms in environmental sex determination system by using the comparative genomics and functional genomics tools.
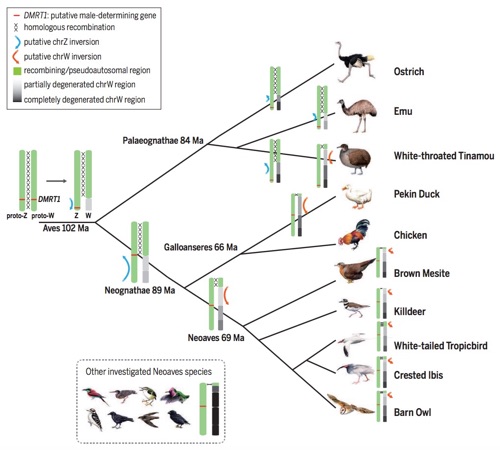
As one of the subprojects in the avian phylogenomic program, we have analyze newly available genomes of 17 bird species representing the avian phylogenetic range to reveal the evolution of sex chromosome in birds (Science 2014). With more completed genome data available at B10K family level, we will continue this study to explore the sex chromosome evolution history across entire avian class. Particularly, we are focusing on the ratite family, which is the basal family of extent bird class, because our previous study has suggested this group of bird species have homomorphic sex chromosomes with extensive recombining and euchromatic PARs that resemble the ancestral state.
- Environmental and epigenetic regulation of sex development
Mechanisms of sex determination can be broadly divided into two major categories: genetic sex determination (GSD) and environmental sex determination (ESD). Generally, the sex of GSD species will be fixed for life, but in some GSD species, the primary sex can be altered during development, such that individuals can develop into the opposite sex without changing their genotype. This phenomenon is known as environmental sex reversal (ESR) and is relatively common in insects, fishes, amphibians, and reptiles. Half-smooth tongue sole, Cynoglossus semilaevis, employs a female heterogametic sex determination system (ZW♀/ZZ♂). However, despite the primary determination of sex by chromosome inheritance, the ZW genetic females are known to be sex-reversed to phenotypic males when reared under normal conditions (22°C). Therefore, this species is an excellent model to investigate the role of DNA methylation in transition from GSD to ESD. Our previous genome study has constructed the origin and evolutionary process of the ZW chromosome system in this fish was derived from the same ancestral vertebrate protochromosome as the avian W and Z chromosomes (Nature Genetics 2014). We compared the gonadal DNA methylomes of pseudomale, female, and normal male fish revealed that genes in the sex determination pathways are the major targets of substantial methylation modification during sexual reversal. Methylation modification in pseudomales is globally inherited in their ZW offspring, which can naturally develop into pseudomales without temperature incubation. This study proved that epigenetic regulation plays multiple crucial roles in sexual reversal of tongue sole fish. We also offer the first clues on the mechanisms behind gene dosage balancing in an organism that undergoes sexual reversal. Finally, we suggest a causal link between the bias sex chromosome assortment in the offspring of a pseudomale family and the transgenerational epigenetic inheritance of sexual reversal in tongue sole fish. To further understand how environmental factor (the high temperature) influences the development of the fish and changes its sex determination pathway, we continue using this fish as model and generate more epigenomic data and histone modification data.
- Functional roles of highly conserved genomic elements in bird development
It has been argued for several decades that the phenotypic variations within and between species can be established by modification of cis-regulatory elements, which can alter the tempo and mode of gene expression. Nevertheless, we still have little knowledge about the genetic basis of macroevolutionary transitions that produced the phenotypic novelties that led to the great leap of evolution and adaptation to new environment. One of these magnificent examples is the transition from non-avian dinosaurs to birds that has created numerous evolutionary innovations such as self-powered flight and its associated wings with flight feathers. Despite extensive paleontological and anatomical studies on birds and their near close relatives, the genetic background for these emerging specializations remains unclear. We hypothesize that the underlying genes and/or their cis-regulatory elements should be maintained by strong selective constraints throughout the avian class, and show distinctive differences from other non-avian species. Based on this hypothesis, we identified millions of avian-specific highly conserved elements (ASHCEs) from the whole genome comparison for the 48 bird species (Nature Communications, 2017). It is worth noting that, 99% of these ASHCEs locate in non-coding regions. Together with the analyses for histone modification profiling data from the chicken embryo and limb buds, we found that the ASHCEs have strong potential to function as cis-regulatory elements.
The above finding inspires us to explore the functional roles of ASHCEs in the development of avian-specific features. Our pilot investigation on the limb buds using comparative gene expression analyses between birds and other vertebrates identified a list of genes showing unique expression patterns in birds. In particular, our experiments offer evidence for a specific role of Sim1 in flight feather development and the essential switch on and off regulatory role of ASHCE on its avian-unique expression pattern. This study demonstrates a framework for future study to identify genomic modifications responsible for various levels of lineage-specific phenome innovations by integrating analytical tools from genomics, developmental biology, and paleontology. We postulate that there are more functionally relevant ASHCEs that coordinate the regulation of additional avian class-specific features, some of which might have been shared with their ancestors among theropod dinosaurs and served as genomic mechanisms for macroevolution. Therefore, we plan to perform the comparison of transcriptome and histone modification data from other tissues during embryos between birds and other vertebrate groups, and to further explore the potential roles of ASHCEs in corresponding to the development of other bird features.